Background
Background
A great deal of time is spent in medicine reading and writing case reports. Essentially, clinical features are listed and a diagnosis made. Excluding those cases that point to a novel means of treatment, a case report is often noteworthy simply because the diagnosis is rare, or because the clinical features were most un-likely to be associated with the diagnosis. This hardly seems a reliable method of archiving medical knowledge.
Much less time is spent on attempting a method of diagnosis that is more systematic than the recalling of case reports. One can see that if one did wish to move medical diagnosis into the information age, natural instinct would be to use an internet search engine to enter a list of clinical features and see what disease diagnoses were associated with these terms. Unfortunately, internet search engines concern themselves only with the popularity of search terms and because of the dominance of case reports such practice may be likely to throw up the least likely cause of those features, or that which is most “titillating” to those who most perform internet searches.
There have been attempts to provide a more balanced means of linking clinical features with diseases and hence making clinical diagnoses. Rare disease with a large number of different clinical features are least easily diagnosed by clinical experience or key investigations, and so the focus of these attempts has been on rare genetic diseases using ever-expanding databases such as Orphanet, Online Mendelian inheritance in Man (OMIM) and the London Dysmorphology Database and the Pictures of Standard Syndromes and Undiagnosed Malformations (POSSUM).
One method of searching for clinical features on these databases is simple text matching. A way of quantifying the match is the feature vector method, which calculates the mathematical overlap between the Query (the clinical features of the case) and the Disease (the clinical features of the disease). A vector of the query is calculated with dimensions for each feature and a value of 1 if present and 0 if absent. The same is done for the disease. The dot product of the two vectors is the strength of the match (a 1 for both query and disease will sway the two vectors in a common direction, and a 0 for both will leave their relationship unchanged, while a 0 and a 1 will make one move away from the other).
A potentially better quantification of matching is to take into account the different specificities of different clinical features. If a clinical feature is present in only a few diseases, its annotation (the linkage of a clinical feature to a disease) is more specific for that disease (in database terms this is called the information content (IC)) and so that linkage should have more weighting. The IC is simply the negative log of the frequency of the annotation. For example, AV block is a term that annotates 3 diseases in the 4813-disease OMIM database. The frequency is 3/4813. Loge of this is -7.38 and minus loge of this is 7.38. A much more general term will have many annotations and a much lower negative log, tending towards zero. The ICs of all the clinical features of the query can be summed or otherwise combined to provide an overall match.
The authors of the presented paper have described a further refinement of this method. This is called the Ontology Similarity Search (OSS). Instead of simply matching the text of terms, they fit clinical features into a standardised language within an ontological framework. This means that the features are related to one another in a hierarchy, with more general terms higher in the hierarchy and more specific subcategories of those general terms lower in the hierarchy. While “parent” terms obviously have many “child” terms, child terms can also belong to multiple parent terms. For example, optic atrophy could be a child of demyelinating disease and also a child of visual disturbance. Their ontology is called the Human Phenotype Ontology (HPO) and has around 9000 terms.
The advantage of using the ontology is that if a clinical feature of a case does not fit the clinical features of the disease, but shares a parent term with one of the features of the disease, instead of scoring a zero match, this scores as a match but less so than if the match was with the specific terms. The method specifically find the most informative common ancestor of the two different clinical features, and uses the IC of that term. Being a more general term, it will be a feature of more diseases and so have a lower IC. (In the database, ancestor terms are implicitly annotated when child terms are annotated.) The overall strength of match is the average of all the ICs – there will always be a IC for each feature, even if it is just that they are both a feature of “any disease”, which of course has an IC of zero and would bring down the average.
Summary of the Paper
The presented paper, Clinical Diagnostics in Human Genetics with Semantic Similarity Searches in Ontologies by Köhler et al. (Am J Hum Genet. 2009 Oct 9; 85(4): 457–464), describes a further refinement of the method using a statistical treatment. For a given disease, if random clinical features from the HPO were selected one would expect a lower OSS score than for a patient who actually had the disease. If the OSS for random features were repeated many times, a distribution would be created and so one could then look at the real patient OSS and determine a p-value on this distribution. If the real OSS was higher than 95% of the random OSS scores, the p-value would be lower than 0.05 and indicate a likely match. Furthermore, if the same features were compared with different diseases and their random OSS distributions, a ranking of the likelihood of diseases could be determined by ranking the corresponding p-values. They call this the OSS –PV.
Since they considered it too onerous to enter, within the framework of the terms of the HPO, the clinical features of real patients with known diseases, they used simulated patients. This was done for 44 diseases, where they created a “patient” having a disease with a selection of the clinical features of the disease weighted by how commonly those features were found in that disease. For each disease 100 patients were created, so if from the clinical literature a feature is found in 1% of cases with the disease, 1 of the 100 simulated patients would have that feature.
They added “noise” to the process by adding to the patients some random features that were not part of the disease, and “imprecision” to the process by replacing some features with their HPO parent terms.
Then they looked at the rank position of the true disease among all the 5000 or so database diseases found by the different methods. The closer the rank position to the true position (first!), the better the method performed.
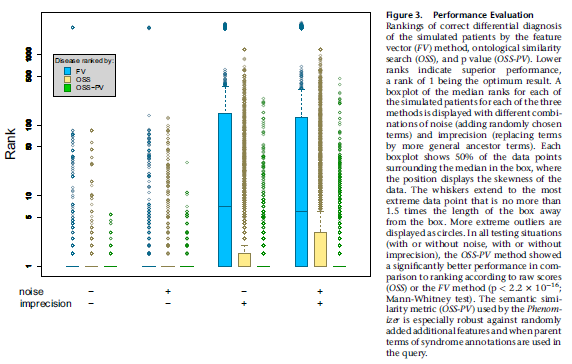
Unsurprisingly, the performance of the feature vector method, as shown by box plots of rankings for all 44 diseases tested, was found to suffer when imprecise terms were used, because that was the point of using the ontological system. The OSS-PV method more modestly outperformed the raw OSS method when noise and imprecision were added.
As the authors point out, the OSS method potentially suffers from the fact that it only matches query terms with disease terms. If a disease also had many terms that did not match the query terms, surely the overall match would be less specific. This can be taken into account by performing a symmetrical similarity search, where the OSS is the average of the matches of the query to the disease and the matches of the disease to the query. However, they did not use this method in their presented data, only stating that when they used it the symmetrical OSS-PV still significantly outperformed the feature-vector method. They do not state that it still outperforms the symmetrical raw OSS.
Another point raised by the paper is that if one finds on a disease search that no disease fits the features with a p-value less than 0.05, exploration could be made of other clinical features, or child features of the entered clinical features that would have a higher information content and provide a more significant match. Going back and looking for a specific feature, or performing a specific investigation, would be an example of this.
Journal Discussion
As described in the introduction, any attempt to quantify and rationalise differential diagnosis should be lauded and this paper clearly describes progressive refinements of this process. It is almost negligent to have all the data available on thousands of diseases and not to use them because the unaided human mind simply cannot store so much information.
However, a number of further refinements and limitations present themselves.
First, the matching of terms is still semantic rather that systematic. While a knowledge-based approach, it nevertheless does not rely on understanding of disease pathophysiologies and pathognomonic features. Some clinical features that share a close parent may in fact best distinguish diseases rather than be considered loosely positively associated features. This may apply particularly in neurology where there is a more systematic approach. For example, upper motoneurone lesion and lower motoneurone lesion may be considered together and share a common parent in “motor neurone lesion”, but apart from the case of motoneurone disease, they split the differential diagnosis more than upper motoneurone lesion and no motor lesion at all. They are semantically similar but nosologically opposite. Horizontal supranuclear gaze palsy and vertical supranuclear gaze palsy may share a strong information content parent, but may be the feature that best separates Gaucher disease from Nieman Pick disease.
This leads to the second point. The frequency, or sensitivity, of a clinical feature in a disease is not considered, although ironically considered when creating the simulated patients with the 44 tested diseases. In large part this reflects the lack of clinical data in the databases themselves. It is regrettable that case reports are not combined into case series which contain information on the frequencies of occurrence of clinical features, or when there are case series, these data are not actually collected systematically. If a clinical feature occurs in 1% of cases of one disease and 100% of cases of another disease, clearly the annotation of the feature for the second disease should be considered far stronger than for the first. Instead, because there are no such data, they are given equal weight; the weighting only considers whether or not the feature is also found in a number of other diseases, not how commonly it is found in those diseases.
There is no consideration of how common the disease is in the first place. While restricting themselves to rare and genetic diseases by definition, there can be a frustrating tendency for searches to throw up the least likely diagnosis. It is often the case in practice that the clinician does not know in advance that the patient has a rare genetic disease, and a diagnostic tool should be most useful to those with least intimate knowledge of the database. Thus, when entering the features dystonia, spastic hemiparesis and spastic dysarthria in a case of cerebral palsy, it comes as a surprise when the top diagnosis is cleft palate-lateral synechia syndrome.
Finally, the methods assume that clinical features are independent. In fact, many clinical features are strongly interdependent; they especially occur together. The association of the second feature is not really very additionally informative if the first is present. This problem would be common to most forms of differential diagnosis calculators, including those using Baysian methods, and could only be solved if there were data on the interdependence of clinical features in different diseases; currently it is hard to find even raw frequency data for most diseases.
The point that the authors raise about using their App to find features that would be more specific in making a diagnosis is an interesting one, and opens a new approach to diagnosis and refinement of the process of often expensive and sometimes risk-associated investigation. One could imagine the improvements in medical care that would arise from use of an App that gave a differential diagnosis based on initial clinical information and then showed the relative power of different investigations in narrowing that differential.
A further use of these methods would be in creating diagnostic criteria. While clinical practice is rightly focused on the most likely diagnosis in a patient, clinical research is focused on a group of patients where the diagnosis is certain, i.e. specificity at the expense of sensitivity. Currently, diagnostic criteria seem to be set largely by “workshops” – gatherings of the great and the good usually in an exotic location who draw up a list of features, create two categories of importance and then decide how many features are required for a “definite diagnosis”. Using a quantified method such as that described in this paper for every study patient and including only patients where the diagnosis reaches a threshold p-value score would seem to be a far more reliable method.
The paper on which this journal club article is based was presented by Dr John McAuley, Consultant and Honorary Senior Lecturer in Neurology at Queens Hospital, Romford.